hankyoreh
Links to other country sites 다른 나라 사이트 링크
S. Korean research team compiles high-resolution genetic map of novel coronavirus
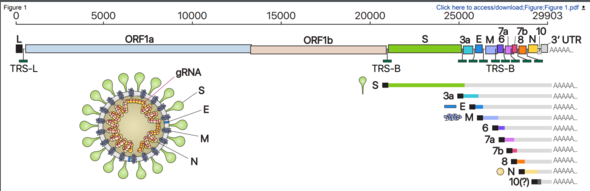
In a joint study with the Korea National Institute of Health, under the Korea Centers for Disease Control and Prevention (KCDC), the Center for RNA Research at the Institute for Basic Science (IBS) has managed to compile a high-resolution genetic map based on an analysis of the transcriptome of the RNA created in host cells by SARS-CoV-2, the novel coronavirus that causes the disease COVID-19. The researchers discovered dozens of previously unknown types of RNA and at least 41 RNA modification sites that present possible targets for medication being developed to treat COVID-19.
A research team led by V. Narry Kim, director of the center, and Chang Hye-shik, professor of biological sciences at Seoul National University, analyzed all the base sequences in the transcriptome (set of all RNA molecules) that SARS-CoV-2 creates in its host cells. Through this analysis, they were able to determine with accuracy the location of the genes in genomic RNA.
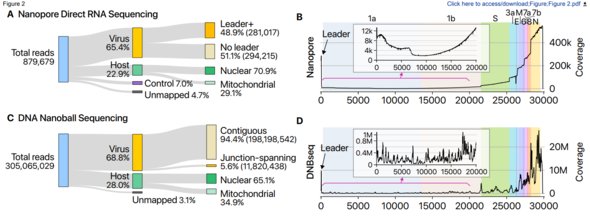
A COVID-19 infection begins when SARS-CoV-2, after infiltrating a host cell, copies its genomic RNA and produces a range of subgenomic RNA. Previous studies had only predicted gene location based on information about genomic RNA.
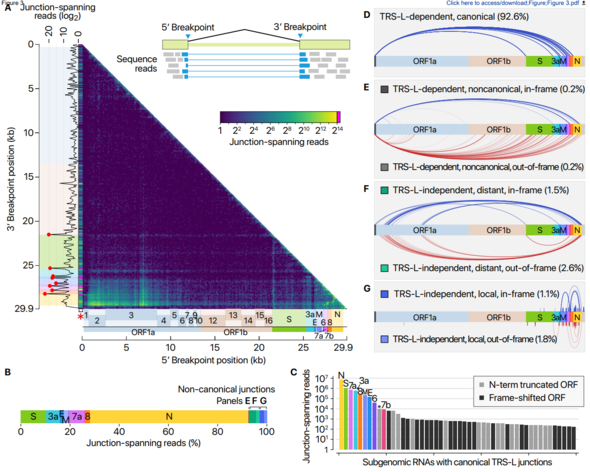
This study confirmed that only nine of the previously predicted ten subgenomes actually exist and identified a variety of chemical modifications that could present new characteristics. This helps researchers understand the coronavirus’s life cycle and pathogenicity, which could be used in developing a new therapeutic strategy.
“The newly discovered RNA and RNA modification sites represent potential targets in the development of viral medications,” V. Narry Kim said.
“Now that we’ve accurately determined the quantity of each transcriptome in SARS-CoV-2, that could serve as the basis for improving the polymerase chain reaction technique used to diagnose the disease,” added Chang Hye-shik.
This study applied the nanopore-based direct RNA sequencing approach (a next-generation technique for analyzing base sequences that was used for the first time in South Korea) to inactivated viruses provided by the KCDC. This new approach makes it possible to analyze SARS-CoV-2’s lengthy RNA sequences directly, without chopping them up. Under the typical approach, RNA is converted to DNA for analysis.
This paper was published online in Cell, an authoritative journal in the field of biology, on Apr. 9.
By Kim Jeong-su, senior staff writer
Please direct comments or questions to [english@hani.co.kr]

Editorial・opinion
![[Column] Tariffs on China: Trump was dumb, Biden dumber [Column] Tariffs on China: Trump was dumb, Biden dumber](https://flexible.img.hani.co.kr/flexible/normal/500/300/imgdb/original/2024/0520/191716191153918.jpg) [Column] Tariffs on China: Trump was dumb, Biden dumber
[Column] Tariffs on China: Trump was dumb, Biden dumber![[Column] What if Seoul took reunification by force off the table? [Column] What if Seoul took reunification by force off the table?](https://flexible.img.hani.co.kr/flexible/normal/500/300/imgdb/original/2024/0520/3017161928630494.jpg) [Column] What if Seoul took reunification by force off the table?
[Column] What if Seoul took reunification by force off the table?- [Editorial] Intensifying US-China rivalry means Seoul must address uncertainty with Beijing sooner than later
- [Column] When ‘fairness’ means hate and violence
- [Editorial] Yoon must stop abusing authority to shield himself from investigation
- [Column] US troop withdrawal from Korea could be the Acheson Line all over
- [Column] How to win back readers who’ve turned to YouTube for news
- [Column] Welcome to the president’s pity party
- [Editorial] Korea must respond firmly to Japan’s attempt to usurp Line
- [Editorial] Transfers of prosecutors investigating Korea’s first lady send chilling message
Most viewed articles
- 1Xi, Putin ‘oppose acts of military intimidation’ against N. Korea by US in joint statement
- 2Kim Jong-un wanted to meet with residents of shelled Yeonpyeong Island in South, Moon recalls in mem
- 3To weigh costs and benefits, Korea must stop treating US troop presence as a sacred cow
- 4[Column] What if Seoul took reunification by force off the table?
- 5Berlin mayor hints at tearing down ‘comfort women’ memorial in city
- 6[Column] Tariffs on China: Trump was dumb, Biden dumber
- 7[Editorial] Transfers of prosecutors investigating Korea’s first lady send chilling message
- 8[Exclusive] Truth commission to seek additional murder charges for figures behind 1980 Gwangju massa
- 9Naver’s union calls for action from government over possible Japanese buyout of Line
- 10[Column] US troop withdrawal from Korea could be the Acheson Line all over